 Evolution
Evolution
 Intelligent Design
Intelligent Design
A New Paper from Biologic Institute: "Enzyme Families — Shared Evolutionary History or Shared Design?"
I am pleased to announce the publication of a new paper from Biologic Institute, a research organization devoted to investigating the limits of unguided evolution and advancing the development of a new paradigm for biology based on intelligent design. This paper, "Enzyme Families — Shared Evolutionary History or Shared Design? A Study of the GABA-Aminotransferase Family," is the closing chapter of our long-term study of bacterial enzymes to determine if they can be coopted to new functions. The answer to this question is important for the evolution debate. If enzymes can’t be recruited to genuinely new functions by unguided means, no matter how similar they are, the evolutionary story is false.
Published in the journal BIO-Complexity, the work was done by Marci Reeves, Doug Axe, and myself.
In a previous paper we described the difficulty of coopting the enzyme Kbl to perform the function of BioF. The two enzymes are very similar in structure (see below) but have different reaction chemistries and different functions in the cell. We wanted to know if a mutated Kbl could replace missing BioF function. After changing nearly every amino acid in Kbl’s active site (where its chemistry is carried out) to look like BioF, Kbl never was able to make the switch to BioF’s function.
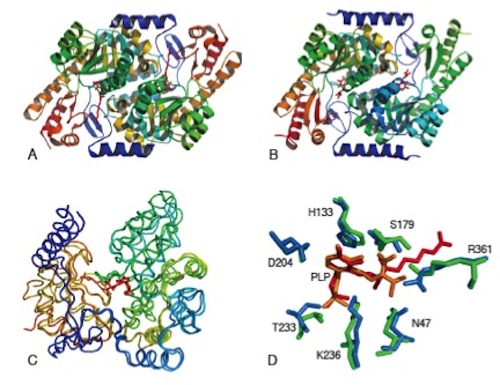
(A) BioF dimer; (B) Kbl dimer; (C) backbones of the two enzymes aligned against each other; (D) the amino acid side chains in the active site(s) of each enzyme that carries out each reaction — BioF is blue and Kbl is green. The red and orange are the reaction products still in the active site(s).
In this paper we expanded the story to include nine of the most closely related enzymes to BioF, including one that is supposed to be able to carry out both BioF’s and Kbl’s chemistry. Using random mutagenesis we tested every single-base mutation in those nine genes. None of them was within one mutation of cooption. We went on to test for cooption the two most likely enzymes by generating two-base combinations of mutations. After testing 70 percent of all possible two-base mutations for each enzyme, or about 40 million cells each, that also failed.
What does this mean? In an evolutionary scenario, to get an enzyme to switch functions the first step is to make a spare copy that can be mutated without destroying a function the cell needs. Second, the cell has to overproduce the mutating enzyme, because any newly emerging enzyme will be very bad at the job at first. To compensate there will need to be lots of enzyme around. Third, there is the problem of finding the right combination of mutations by random search.
Taken together, since we found no enzyme that was within one mutation of cooption, the total number of mutations needed is at least four: one for duplication, one for over-production, and two or more single base changes. The waiting time required to achieve four mutations is 1015 years. That’s longer than the age of the universe. The real waiting time is likely to be much greater, since the two most likely candidate enzymes failed to be coopted by double mutations.
We have now addressed two objections raised by our critics: that we didn’t test the right mutation(s), and that we didn’t use the right starting point. We tested all possible single base changes in nine different enzymes, Those nine enzymes are the most structurally similar of BioF’s entire family We also tested 70 percent of double mutations in the two closest enzymes of those nine.
Finally, some have said we should have used the ancestral enzyme as our starting point, because they believe modern enzymes are somehow different from ancient ones. Why do they think that? It’s because modern enzymes can’t be coopted to anything except trivial changes in function. In other words, they don’t evolve!
That is precisely the point we are making.
