 Intelligent Design
Intelligent Design
Some Perspective on the “Mechanical Prowess” of DNA Replication
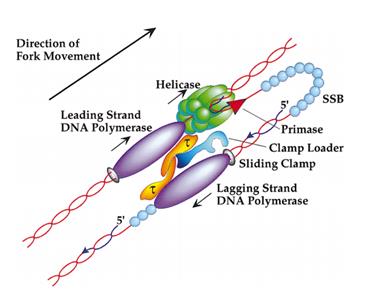
I recently published two articles (here and here) on the mechanisms by which a cell’s DNA is replicated. Now I’ve just encountered a paper, published in the journal Cell in 1998, that does an excellent job of describing the engineering prowess of these molecular machines. The paper is titled “Polymerases and the Replisome: Machines within Machines,” and it makes the following statements about the processes of DNA replication:
Synthesis of all genomic DNA involves the highly coordinated action of multiple polypeptides. These proteins assemble two new DNA chains at a remarkable pace, approaching 1000 nucleotides (nt) per second in E. coli. If the DNA duplex were 1 m in diameter, then the following statements would roughly describe E. coli replication. The fork would move at approximately 600km/hr (375 mph), and the replication machinery would be about the size of a FedEx delivery truck. Replicating the E. coli genome would be a 40 min, 400 km (250 mile) trip for two such machines, which would, on average make an error only once every 170 km (106 miles). The mechanical prowess of this complex is even more impressive given that it synthesizes two chains simultaneously as it moves. Although one strand is synthesized in the same direction as the fork is moving, the other chain (the lagging strand) is synthesized in a piecemeal fashion (as Okazaki fragments) and in the opposite direction of overall fork movement. As a result, about once a second one delivery person (i.e. polymerase active site) associated with the truck must take a detour, coming off and then rejoining its template DNA strand, to synthesize the 0.2km (0.13 mile) fragments.
It can hardly be disputed that DNA replication is extraordinary. Stay tuned for a forthcoming article on DNA helicase.
