 Evolution
Evolution
What Really Happened at Wistar II
In June 2007, I attended a private conference in Boston, commemorating the famous Wistar Symposium of 1966. All participants were asked to keep the proceedings confidential — and all did. One participant, however, Daniel Brooks of the University of Toronto, later violated that agreement, and published his version of events on-line.
 That account can be found here.
That account can be found here.
Several times now, that erroneous account has been quoted against me in different venues. In the interests of truth, I am therefore setting the record straight. This exercise may shed some light on the way science is done.
The research I described
At the meeting I presented preliminary work that later became the paper on enzyme evolution published last year in the peer-reviewed journal BIO-Complexity. The work involved screening cells for their ability to grow on medium lacking biotin, an essential vitamin. Normally bacterial cells can either take up biotin from their environment, or make it themselves using a four-step dedicated pathway. The cells I worked with lacked the first enzyme in that pathway, called BioF, and thus could not make biotin. We were testing whether a structurally similar enzyme called Kbl could be mutated enough to allow it to substitute for the missing BioF, and thus allow the cells to grow.
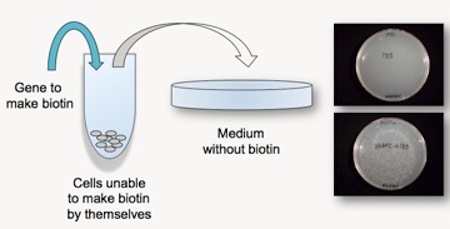
Cells normally only need as little as 10 molecules of biotin in order to grow, so the growth assay was very sensitive to exogenous biotin. But it was also sensitive to cell density, presumably because crowded cells could scavenge biotin from their dying neighbors when on minimal medium.
I normally plated cells at a low density of 500 colonies or less per plate. But one experiment was plated at about 5000 colonies per plate, and left to grow over the weekend. When I checked the plates, I found that a few colonies had grown where none were expected. I isolated those colonies and checked their ability to grow on minimal medium again. It was reproducible. I was excited, because I thought I might have found a mutant strain that could now make biotin, either using Kbl or some other bypass mechanism to replace the missing BioF.
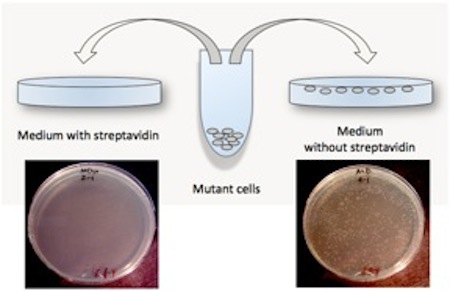
I verified that the cells still lacked BioF. So far so good. But then it occurred to me that they had been isolated from crowded plates, and that they were slow to grow. So I performed one more experiment to see if they were making biotin, or simply scavenging it from their medium more effectively. I plated the cells to minimal medium with and without streptavidin added. Streptavidin sops up any biotin in the medium, binding it very tightly, but does not otherwise effect cell growth.
The mutant strain I had found that could grow slowly on minimal medium could not grow on minimal medium plus streptavidin at all. Thus the mutation these cells carried, whatever it was, did not allow them to make biotin for themselves. It only allowed them to take up biotin from their environment more readily.
How my description, and the reaction to it, were distorted
This is the result I reported at the private meeting. No mistake — the mutated cells grew on medium without added biotin, but this ability was the result of increasing their ability to scavenge biotin from the medium, not because they could make biotin. They had not found a way to replace BioF function, which was the whole point of the experiment.
I explained this to the group, and one participant, biologist G�nther Wagner of Yale, said, smiling, “So, a beneficial mutation happened right in your lab?” I said “Yes,” also smiling, and everyone laughed. Then the session ended.
G�nter Wagner (and most of the other participants, I assume) knew that the mutant strain I found did not solve the problem I had posed — namely, how new enzyme functions, or the ability to make biotin, arise in the first place. It was beneficial only in the sense that the mutant cells could scavenge biotin from the medium better than before, which allowed them to grow better. In fact, he probably realized that it demonstrated just how sensitive my assay was. If the cells could find a way to make or find even a little biotin, it would have been enough for them to grow.
In his leaked report, Dan Brooks claimed that I had found a mutant with beneficial new information. This is stretching things quite a bit. What I probably found was a mutant with an extra copy of the genes required to transport biotin from the medium, or with a higher affinity for biotin. That kind of mutation would help glean every available biotin molecule, but it would not help make biotin in the first place. There was no new genetic function generated, or sufficient information to make biotin. In the end, this adaptation would be a dead end for these cells once exogenous biotin was exhausted.
In charity, I would offer that perhaps Brooks didn’t follow the details of the experiments I discussed because he is not a geneticist or a molecular biologist. The alternative explanation, that he deliberately misrepresented things to put me in a bad light and to score a talking point, would be unfortunate.
Brooks’s story has been taken up by Internet gadflies. Some have implied that I suppressed this result because it did not fit my agenda, and disproved my own thesis. They have accused me of dishonesty. Now that this true account is available, however, critics who continue to spread the false account will have no excuse.
Biologic Institute is in the business of doing science. We are not interested in point-scoring, name-calling, or Internet flame wars. We will continue to do independent, careful science, and to publish our work in peer-reviewed settings for all to read and evaluate. Those who are interested can continue to follow us here at Evolution News & Views and at Biologic Perspectives.
Image: Boston skyline, afagen/Flickr.

