 Evolution
Evolution
The Challenge to Darwinism from a Single Remarkably Complex Enzyme
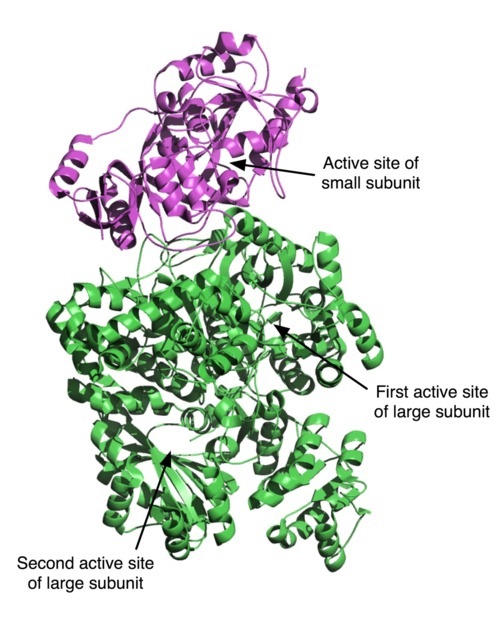
Meet carbamoyl phosphate synthetase (CPS), a remarkably complex enzyme. This enzyme uses bicarbonate, glutamine, ATP, and water to make carbamoyl phosphate via a multi-step reaction at three separate active sites, involving several unstable intermediates. See here for details.
CPS is made of two protein chains with a combined length of over 1,400 amino acid residues. We now know from extensive biochemical data that a fully coupled CPS requires the hydrolysis of one glutamine and two molecules of MgATP for every molecule of carbamoyl phosphate formed. The three active sites of the enzyme maintain the overall stoichiometry of the reaction, without wasteful hydrolysis of glutamine and/or MgATP. In order to couple the reactions efficiently, the enzyme uses internal molecular tunnels for sequestering and rapid transfer of reactants between active sites, and allosteric conformational changes to synchronize their activity. This is remarkable, given that the first and last active sites are separated by nearly 100 �. But even more remarkable is the fact that this enzyme carries out a series of reactions involving unstable intermediates with a half-life of seconds to milliseconds.
How does a neo-Darwinian process evolve an enzyme like this? Even if enzymes that carried out the various partial reactions could have evolved separately, the coordination and combining of those domains into one huge enzyme is a feat of engineering beyond anything we can do.
For more information about the challenge to Darwinism presented by protein folds, go here.
Cross-posted at Biologic Perspectives.
